
RdRp Collaborative Analysis Tool with Collections of pHMMs
RdRpCATCH - the RdRp Collaborative Analysis Tool with Collections of pHMMs represents a collaborative effort to consolidate various publicly available RdRp pHMM resources into a single, user-friendly platform. RdRpCATCH allows for seamless scanning of metatranscriptomics assemblies for RNA virus discovery and provides follow-up taxonomic annotation of the detected contigs.
How It Works
Complete workflow overview
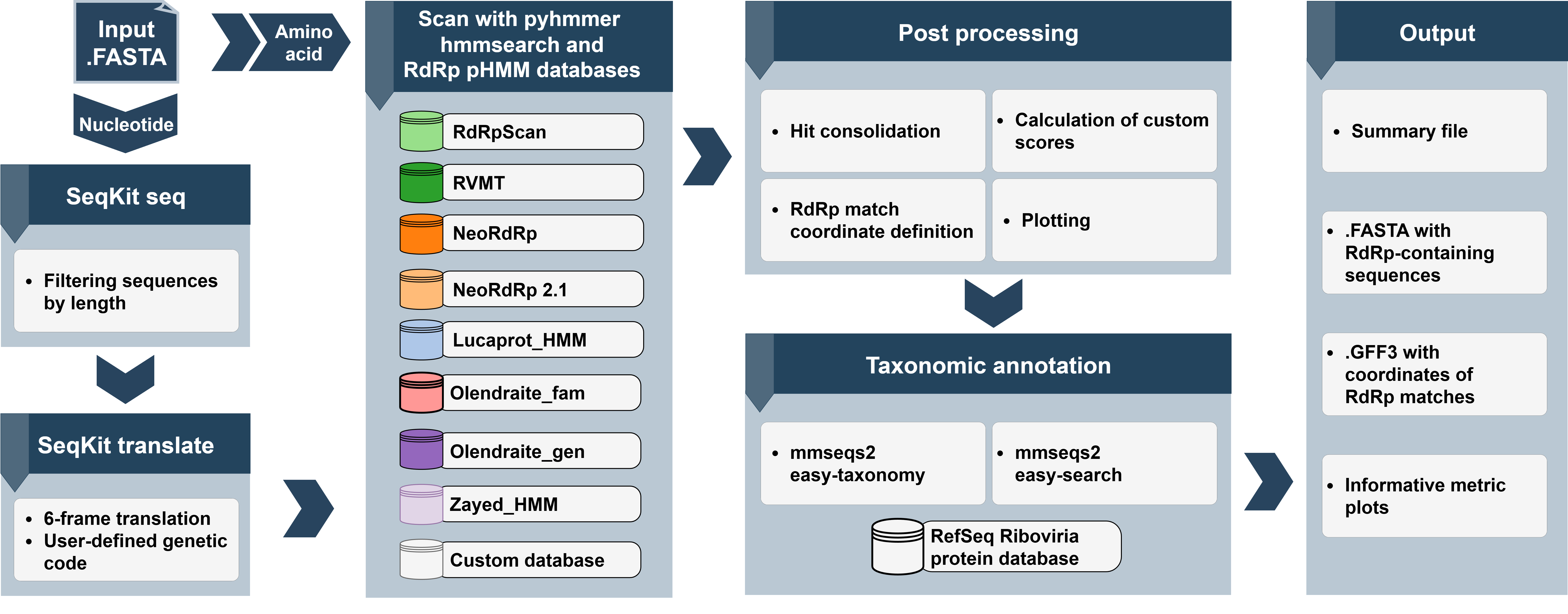
Figure 1: RdRpCATCH workflow overview showing the complete analysis pipeline
Key Features
Discover the key features of RdRpCATCH
Comprehensive Databases
Access and use multiple curated RdRp pHMM databases specialized for comprehensive RNA virus detection.
Easy Upload
Simple drag-and-drop interface for FASTA files. Supports both nucleotide and protein sequences.
Coverage metrics
Calculation of contig and profile coverage offer valuable insights into the RdRp completeness.
Taxonomic Annotation
Automatic taxonomic classification of detected contigs using an NCBI RefSeq Riboviria protein database and mmseqs2.
Comprehensive Results
Download detailed results including annotated sequences, taxonomic classifications, and visualization plots for further analysis.
Secure & Reliable
Your data is processed securely with automatic cleanup. Results are available for 7 days with secure access links.
Resources & Downloads
Access additional resources and command-line tools
GitHub Repository
Access the complete source code, documentation, and command-line version of RdRpCATCH for advanced users and developers.
View RepositoryZenodo Archive
Download the complete tool package including databases, scripts, and documentation for offline use.
Download PackageRdRp Summit Community
RdRpCATCH emerged as a collaborative project from the global RdRp Summit community. Join the community at rdrp.io for more information about the RdRp Summit, Slack, and Journal Club.
Visit rdrp.ioCitation
If you use RdRpCATCH in your research, please cite RdRpCATCH and the underlying databases:
Karapliafis, D., Neri, U., Olendraite, I., Charon, J., Sakaguchi, S., Hou, X., De Ridder, D., Zwart, M. P., & Kupczok, A. (2026). RdRpCATCH: A unified resource for RNA virus discovery using viral RNA-dependent RNA polymerase profile Hidden Markov models. BioRxiv. https://doi.org/10.64898/2026.02.05.703936
Our platform supports the following databases:
1. Lucaprot_HMM: Hou, X. and He, Y. et al. (2024) 'Using artificial intelligence to document the hidden RNA virosphere', Cell, 187(24).
doi:10.1016/j.cell.2024.09.027
2. NeoRdRp: Sakaguchi, S. et al. (2022) 'NeoRdRp: A comprehensive dataset for identifying RNA-dependent RNA polymerases of various RNA viruses from metatranscriptomic data', Microbes and Environments, 37(3).
doi:10.1264/jsme2.me22001
3. Neordrp2: Sakaguchi, S., Nakano, T. and Nakagawa, S. (2024) 'Neordrp2 with improved seed data, annotations, and scoring', Frontiers in Virology, 4.
doi:10.3389/fviro.2024.1378695
4. RDRP-Scan: Charon, J. et al. (2022) 'RDRP-Scan: A bioinformatic resource to identify and annotate divergent RNA viruses in metagenomic sequence data', Virus Evolution, 8(2).
doi:10.1093/ve/veac082
5. RVMT: Neri, U. et al. (2022) 'Expansion of the global RNA virome reveals diverse clades of bacteriophages', Cell, 185(21).
doi:10.1016/j.cell.2022.08.023
6. Olendraite_fam: Olendraite, I. (2021) 'Mining diverse and novel RNA viruses in transcriptomic datasets', Apollo.
Link
7. Olendraite_gen: Olendraite, I., Brown, K. and Firth, A.E. (2023) 'Identification of RNA virus–derived rdrp sequences in publicly available transcriptomic data sets', Molecular Biology and Evolution, 40(4).
doi:10.1093/molbev/msad060
8. Zayed_HMM: Zayed, A. A., et al. (2022) 'Cryptic and abundant marine viruses at the evolutionary origins of Earth’s RNA virome.' Science, 376(6589), 156–162.
doi:10.1126/science.abm5847